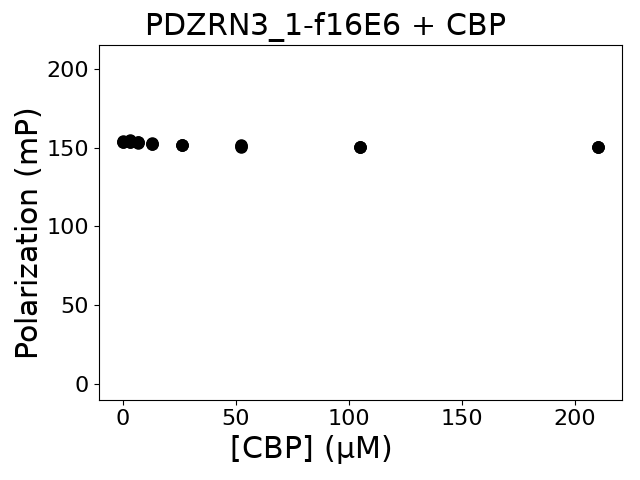
Measured dissociation constant (10-6M):
Standard deviation of dissociation constant (10-6M): 0
Average holdup BI: -0.01
Normalized holdup BI: -0.01
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1