Details of experiments between Q15418 and Q96QZ7
Experiment series 1 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.46
Holdup BI standard deviation: 0.05
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 5
Measured pK d : 4.47
Experiment series 2 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.22
Normalized holdup BI: 0.25
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 4.3
Experiment series 3 Protein A Protein: RPS6KA1 726-735 PBM RVRKLPSTTL Biotin-ado-ado-RVRKLPSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.21
Normalized holdup BI: 0.23
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 4.24
Experimental Details Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
Experiment series 4 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
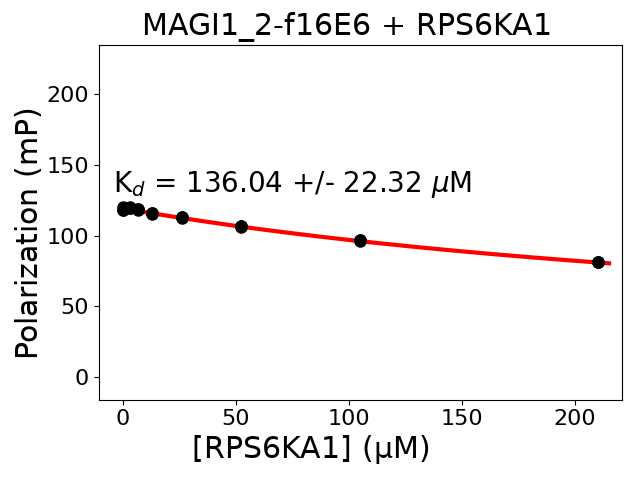
Measured dissociation constant (10-6 M): 136.04
Standard deviation of dissociation constant (10-6 M): 22.32
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit
Number of measurements: 3
Measured pK d : 3.87
Experiment series 5 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.07
Normalized holdup BI: 0.07
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 3.64
Experiment series 6 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.01
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 7 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 1146-1238 PDZ_6 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.02
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 8 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL Biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M): 0
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: SPR
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 9 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 638-725 PDZ_3 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.05
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 10 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 836-926 PDZ_4 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.04
Holdup BI standard deviation: 0.04
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 11 Protein A Protein: RPS6KA1 725-735 PBM RVRKLPSTTL biotin-ttds-RRVRKLPSTTL
Protein B Protein: MAGI1 986-1098 PDZ_5 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.02
Holdup BI standard deviation: 0.05
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 31
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 4
The affinity was below the detection threshold of the assay.
Experiment series 12 Protein A Protein: RPS6KA1 726-735 PBM RVRKLPSTTL Biotin-ado-ado-RVRKLPSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.13
Normalized holdup BI: -0.13
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experimental Details Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
Experiment series 13 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.12
Normalized holdup BI: 0.14
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 15
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 4.05
Experimental Details not reported internal standards of holdup layout #5
Experiment series 14 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
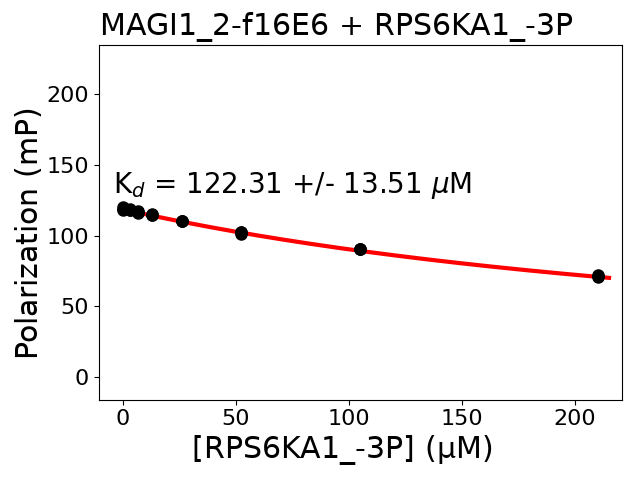
Measured dissociation constant (10-6 M): 122.31
Standard deviation of dissociation constant (10-6 M): 13.51
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit
Number of measurements: 3
Measured pK d : 3.91
Experiment series 15 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.12
Holdup BI standard deviation: 0.12
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 5
Measured pK d : 3.52
Experiment series 16 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0
Normalized holdup BI: -0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 15
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 17 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.03
Normalized holdup BI: 0.03
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 15
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experimental Details not reported internal standards of holdup layout #5
Experiment series 18 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.01
Holdup BI standard deviation: 0.05
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 19 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 1146-1238 PDZ_6 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.04
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 20 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.04
Normalized holdup BI: 0.04
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 15
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 21 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL Biotin-ttds-RRVRKLP(pS)TTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M): 0
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: SPR
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 22 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 638-725 PDZ_3 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.02
Holdup BI standard deviation: 0.01
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 23 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 836-926 PDZ_4 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.03
Holdup BI standard deviation: 0.06
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 2
The affinity was below the detection threshold of the assay.
Experiment series 24 Protein A Protein: RPS6KA1 725-735 PBM (SEP)732 RVRKLPSTTL biotin-ttds-RRVRKLPpSTTL
Protein B Protein: MAGI1 986-1098 PDZ_5 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.01
Holdup BI standard deviation: 0.03
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 40
P value: 0
Experiment method: CALIP_holdup
Details of affinity fitting: BSA and lysozyme internal standards + bacterial cell lysate + normal layout
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 25 Protein A Protein: RPS6KA1 725-735 PBM (TPO)733 RVRKLPSTTL biotin-ttds-RRVRKLPSpTTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0.08
Normalized holdup BI: -0.08
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
Experiment series 26 Protein A Protein: RPS6KA1 725-735 PBM (TPO)733 RVRKLPSTTL biotin-ttds-RRVRKLPSpTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: -0
Normalized holdup BI: -0.01
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 18
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
The affinity was below the detection threshold of the assay.
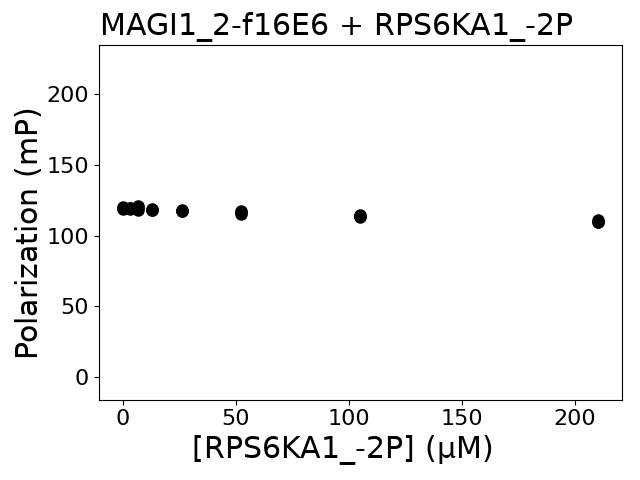
Experiment series 27 Protein A Protein: RPS6KA1 725-735 PBM (TPO)733 RVRKLPSTTL biotin-ttds-RRVRKLPSpTTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M): 0
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit
Number of measurements: 3
The affinity was below the detection threshold of the assay.
Experiment series 28 Protein A Protein: RPS6KA1 725-735 PBM (TPO)734 RVRKLPSTTL Biotin-ttds-RRVRKLPSTpTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.28
Normalized holdup BI: 0.33
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 25
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 4.32
Experiment series 29 Protein A Protein: RPS6KA1 725-735 PBM (TPO)734 RVRKLPSTTL Biotin-ttds-RRVRKLPSTpTL
Protein B Protein: MAGI1 456-580 PDZ_2 his6-MBP-TEVsite-PDZ
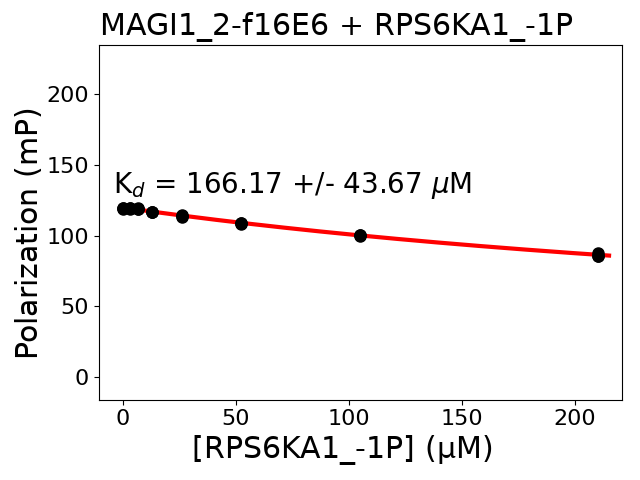
Measured dissociation constant (10-6 M): 71.89
Standard deviation of dissociation constant (10-6 M): 24.9
Average holdup BI: 0
Normalized holdup BI: 0
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 0
P value: 0
Experiment method: competitive FP
Details of affinity fitting: Competitive binding equation in ProFit
Number of measurements: 3
Measured pK d : 4.14
Experiment series 30 Protein A Protein: RPS6KA1 725-735 PBM (TPO)734 RVRKLPSTTL Biotin-ttds-RRVRKLPSTpTL
Protein B Protein: MAGI1 1-106 PDZ_1 his6-MBP-TEVsite-PDZ
Measured dissociation constant (10-6 M):
Standard deviation of dissociation constant (10-6 M): 0
Average holdup BI: 0.05
Normalized holdup BI: 0.05
Normalized holdup BI standard deviation: 0
Immobilized partner concentration (10-6 M): 25
P value: 0
Experiment method: DAPF_holdup
Details of affinity fitting: fluorescein + mCherry internal standards + normalization based on CALIP_holdup data + reverse layout
Number of measurements: 1
Measured pK d : 3.33
Please cite ProfAff! (Gogl, G. et al., Nature Communications 2022) research team of Gergo Gogl .