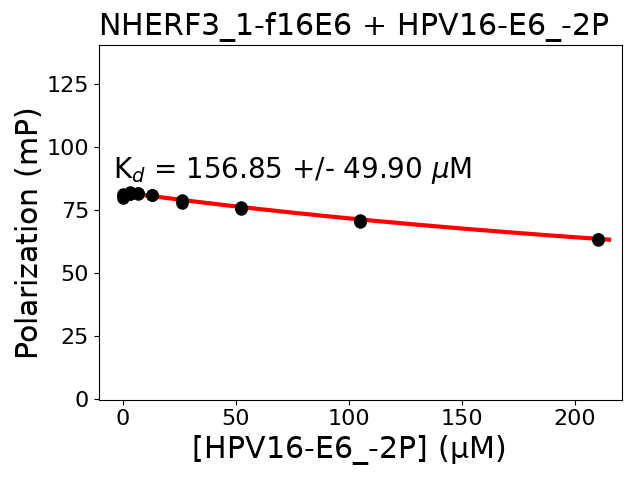
Details of experiments between P03126 and Q5T2W1
Note that the results of all experiments are listed, regardless of the modification states of the fragments.
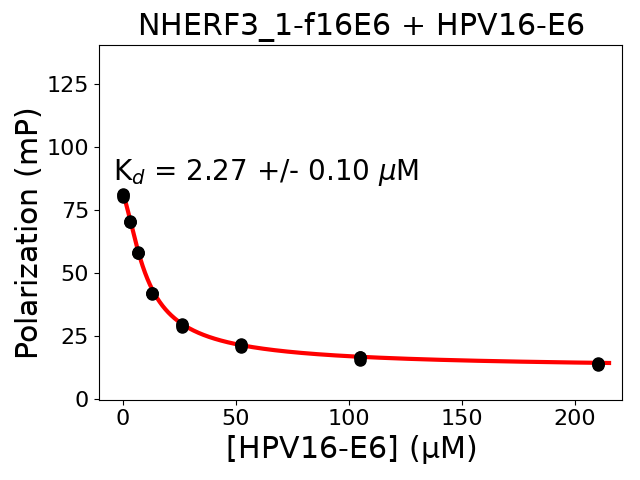
Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
not reported internal standards of holdup layout #5
Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
not reported internal standards of holdup layout #5
Crude peptide synthesis product. The peptide concentration in the affinity conversion step may differ from reality. Not reported internal standards of holdup layout #6
not reported internal standards of holdup layout #5